Top Links
Journal of Microbiology and Modern Techniques
ISSN: 2575-5498
Review on MALDI TOF MS: Modern Disease Diagnosis Approaches in Microbiology and its Mechanisms
Copyright: © Abdi Ahmed Umer. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Related article at Pubmed, Google Scholar
Matrix-assisted laser desorption/ionization time of flight mass spectrometry (MALDI-TOF MS) has become a widely used technique for the rapid, accurate, durable, economical, and trustworthy method for identification of bacteria, mycobacteria and fungal pathogens in the microbiology laboratory. In the last ten years, bacterial identification in microbiology labs has seen major modifications, moving from sluggish conventional microbial identification to quick MALDI TOF MS detection. MALDI has a very simple approach designed for protein detection and reliable technology for precise microbiological identification from culture. Despite the possibility of protein spectrum fluctuation brought about by various MALDI TOF MS procedures. The quickest identification has the most potential to enable for early treatment decision-making
MALDI TOF MS technology can produce mass spectra from samples and has added an advantage of not requiring a predetermined target (in contrast to PCR-based methods of identification). However, the microbe still needs to be cultured, and it cannot handle mixed cultures. The quality and taxonomic range of the reference database are also crucial since only if the microorganism in issue is included in the database can a correct identification be made. MALDI-TOF MS is that it is a highly sensitive technique, capable of identifying even low levels of microbial contamination in samples. This makes it particularly useful in detecting infections and monitoring the spread of infectious diseases.
Keywords: MALDI TOF MS; Modern; Disease; Diagnosis; Microbiology
The recent advancement of Matrix-assisted Laser Desorption/Ionization-Time of Flight Mass Spectrometry (MALDI-TOF MS), has alter the way laboratories method in identify bacteria and fungus. The development of new instruments, along with new data analysis, processing, integration and visualization tools as well as data sharing resources, has accelerated mass spectrometry (MS) based research [1]. MALDI-TOF MS has the benefit of being able to quickly and easily identify bacteria and fungus from colonies produced on culture plates. There are initiatives underway to expand the application of MALDI-TOF MS beyond the microbiological identification of cultured isolates to include bacterial identification directly from clinical samples, determining antibiotics sensitivity and deliver fast data . Conventional biochemical profiling and phenotypic analytical methods has been rapidly replaced by this in a certain diagnostic, research and clinical microbiology laboratories. In addition MALDI TOF MS technology can produce mass spectra from samples and has added an advantage of not requiring a predetermined target (in contrast to PCR-based methods of identification). However, the microbe still needs to be cultured, and it cannot handle mixed cultures. identification of microbes such as bacteria, fungi and yeast. A proteomic fingerprint of an unidentified pathogen is produced, and its genus and species are most likely identified after comparison with a database of reference spectra. The reproducibility of this method is made possible by the analysis of continually produced, extremely abundant proteins, usually ribosomal proteins, in the mass range of 2000 to 20000 Da. MALDI-TOF MS is a ‘soft ionization’ technique, which gently ionizes particles and allows the analysis of larger biomolecules with a time-of-flight analyzer to generate a mass spectrum.
The general specifications of the two major MS-based microbial identification systems available in the global market Biotyper and VITEK MS Plus. There are a number of commercially available methods that usually make a proprietary reference database available with the mass spectrometer, although in house databases (Library) can also be developed. The two main steps in the MALDITOF MS process for identifying microorganisms are spectrum acquisition and spectrum analysis. MALDI-TOF Mass spectrometry is a diagnostic method that uses bacterial protein to analyze the samples' mass-to-charge ratios. Sample preparation is straightforward and repeatable,despite the initial expense of the equipment. MALDI TOF MS technology is more efficient than current phenotypic testing methods in terms of speed, robustness, customization for laboratory demands, and, perhaps most significantly, ease of use. Moreover, compared to other approaches in traditional molecular biology and microbiology, the cost analysis per sample still stays significantly lower . An overview of the MALDI-TOF MS in research and diagnostic microbiology laboratories is given in this manuscript, along with information relates to sample preparation, MALDI TOF MS mechanisms and the expediting of microbiological identification and diagnosis.
For many years, mass spectrometry (MS) technology has been employed in chemistry. Given that varied and distinctive mass spectra were generated from bacterial extracts of various genera and species, Anhalt and Fenselau proposed the use of this method for bacterial characterization in 1975 .Bacterial profiling became possible in the 1980s because of the development of desorption/ionization methods (laser desorption, and rapid atom bombardment) that enable the creation of molecular biomarker ions from microorganisms. The development of soft ionization methods by Tanaka and Fenn in the late 1980s, including matrix-assisted laser desorption/ionization (MALDI) and electrospray ionization, allowed for the investigation of big macromolecules like entire proteins. Two German scientists called Hillenkamp and Karas invented the soft ionization method in 1985.:
Early in the 1990s, the first matrix assisted laser desorption ionization (MALDI-MS) equipment became commercially accessible as a consequence of improvements made to the technology by Karas and Hillenkamp based on the idea of soft ionization [9]. In 1996, it was possible to get MALDI spectral fingerprints from entire bacterial cells without pretreatment prior to the MS analysis [3].Time of flight (ToF) technology was created in the late 1950s, four decades before MALDI. With the use of a nitrogen laser for ionization, Koichi Tanaka and his Japanese colleagues were able to gently ionize considerably larger molecules with success. Later, in 2002, Tanaka received a quarter of the Nobel Prize in Chemistry for his work on the creation of the soft desorption ionization
method for the mass spectrometric characterization of biological macromolecules [4]. This a method rely on forcing charged ions to fly from a source to a detector so that they may be recognized based on the time of their journey to the ion detector. TOF, however, was limited by its low resolution and thus unable to be used in MS methods. When the specimen is being analyzed in the reflection mode, the use of ToF technology in particular is crucial. As a quick, precise, and sensitive method for characterizing and identifying bacteria and fungus, (MALDI-TOF MS) has recently come into use.
For direct samples preparation smear bacterial colony onto each target plate position and allow to dry. Deposit 1 µL of the matrix solution onto each sample spot and allow to dry [5]. Samples are prepared by mixing the analyte with a matrix made of small acid molecules that possesses a strong optical absorption in the range of the wavelength used by the laser device; CHCA α-cyano-4- hydroxycinnamic acid are optimal matrices for the detection of lower mass ions. After co-crystallization of the sample and matrix, the latter absorbs energy from the laser, leading to the desorption and ionization of the analytes in the gas phase. Ions are then accelerated through an electrostatic field (created by a potential of about 20 kV) into the high vacuum flight tube until they reach the detector, with smaller ions traveling faster than larger ones. Thus, the time of flight (TOF) required to reach the detector is dependent on the mass and charge of the bioanalyte, resulting in a spectral profile unique for a given species, composed of peaks ranging usually from 2 to 20 kDa.
Very conserved proteins with housekeeping functions comprise the MALDI-TOF spectra. Detected biomolecules correspond mostly to ribosomal proteins that are abundant, basic, and of medium hydrophobicity, all biochemical traits that favor efficient ionization. Among the proteins ionized during the MALDI process are structural proteins, DNA or RNA binding proteins ribosome modulation factors, carbon storage regulators, cold- shock proteins, and translation initiation factors. These biomarkers generate spectral fingerprints that vary between microorganisms and have peaks specific to genus and species.
For microorganism identification, protein mass patterns are compared within a few minutes with commercially available reference databases [5] for Micro flex LT spectrometer and Vitek Daltonics for Microflex LT spectrometer and Vitek MS for bioMérieux spectrometer, the latest developed by Shimadzu-Biotech, which include species-specific fingerprints of several bacterial and yeast isolates [6]. Through, mass peaks spectra are matched with reference spectra included in the database; this a numerical value is generated that allows accurate and rapid identification of the microorganisms [7].
MALDI-TOF mass spectrometer is composed of three principal units. The first is the ion source that makes ionization and transfers sample molecule ions into a gas phase. The second unit is the mass analyzer that allows ion separation according to mass to charge ratio m/z. The last unit is a detection device for monitoring separated ions [8]. Samples are prepared by mixing the analyte with a matrix made of small acid molecules that possesses a strong optical absorption in the range of the wavelength used by the laser device; CHCA α-cyano-4- hydroxycinnamic acid are optimal matrices for the detection of lower mass ions [9].
After co-crystallization of the sample and matrix, the latter absorbs energy from the laser, leading to the desorption and ionization of the analytes in the gas phase. Ions are then accelerated through an electrostatic field into the high vacuum flight tube until they reach the detector, with smaller ions traveling faster than larger ones. Thus, the time of flight (TOF) required to reach the detector is dependent on the mass and charge of the bioanalyte, resulting in a spectral profile unique for a given species, composed of peaks ranging. Very conserved proteins with housekeeping functions comprise the MALDI-TOF spectra [3].
In an MS analysis utilizing MALDI as a soft ionization mechanism, a saturated solution of a low-mass organic compound, called a matrix, is added to the sample, and the mixture is then spotted onto a metal target plate for analysis. Once appropriately processed samples are added to the MALDI plate, overlaid with matrix, and dried, the sample is bombarded by the laser. This bombardment results in the sublimation and ionization of both the sample and matrix. These generated ions are separated based on their mass-- to-charge ratio via a TOF tube, and a spectral representation of these ions is generated and analyzed by the MS software, generating an MS profile. This profile is subsequently compared to a database of reference MS spectra and matched to either identical or the most related spectra contained in the database, generating an identification for bacteria contained within the sample [10].
Following laser bombardment, ions generated from both the matrix and samples must be analyzed to determine their respective masses and identities. The mass analyzer is the component of a mass spectrometer that functions to determine these representative masses, aiding in the identification of the proteins being analyzed. A variety of mass analyzers exist for measuring ionized proteins from biological samples. In principle, no single analyzer is ideal for all applications, and instruments must be selected on the basis of investigational necessities. In the case of microbial diagnostics, commercial systems have been developed for MALDI-TOF MSbased identification, but it is still necessary to perform instrument calibration and quality control. MALDI utilizes a pulsed ionization source, where a pulse of ions from the specimen is produced by an instantaneous exposure to the laser beam. The pulsed nature of the MALDI process pairs naturally with the TOF mass analyzer, which requires that all ions enter the flight tube simultaneously [11] . Additionally, the TOF mass analyzer is ideal for MALDI, due to its virtually unlimited mass range, which is advantageous because MALDI typically produces singly charged molecular ions that can have a high mass-to-charge (m/z) ratio. The implementation of mass spectrometric techniques into the microbiology laboratory is highly dependent upon method standardization and reproducibility; therefore, mass analyzers are often preselected, optimized, and marketed as part of an instrument package dedicated to microbial identification.
The time of flight (TOF) analyzer is dependent upon the principle that applying an electrostatic field to the ionized samples generated ion with a charge (z) to accelerate, imparting to it some amount of kinetic energy (KE). The ions then move into a field-free drift region, where the only force affecting ionic movement is the kinetic energy from the acceleration Drift time is directly proportional to the m/z ratio. Larger ions will have a longer drift time and smaller molecules will have a shorter drift time, separation of molecules based on mass. This allows for separation of ions originating from samples based on the m/z ratio.
Mass discrimination is one of the frequently reported limitations of MALDI-ToF-MS. This technique generally provides information for relatively lower molecular mass samples. This phenomenon occurs as a result of several different factors such as sample preparation, desorption/ionization procedure and detection. Selecting suitable matrix and ionizing agent are expected to minimize the contribution of sample preparation in mass discrimination. In general, all the elements involved in sample preparation should serve for achieving one goal: highest level of homogeneousity. With regards to the instrumental parameters, many different factors can be responsible for mass discrimination. For instance, the channel plate detector with its limited dynamic range might become saturated by low molecular weight ions such as matrix ions, analyte ions of the shorter chains. For that reason, the higher mass ions are not able to reach to the detector before the smaller molecules that already occupied the detector.
Soft ionization of the analyte molecules occurs by laser irradiation generated ions that can be matrix ions, analyte ions or possibly ionized fragments of the analyte molecules are accelerated and led towards the drift zone. This region is much larger in comparison to the ionization or acceleration zone. At the end of the flight path, the ion detector is located that can register the flight time and the intensity of the individual ions that arrive to the ion detector. As it is expected, heavier analyte ions take longer time to reach to the detector while lighter ions travel the same distance in a shorter period of time. Based on the recorded information by the instrument, the ratio of mass per charge (m/z), m is the mass of the ionized molecule and z is the number of the electrons that have been removed from the molecule. E is the accelerating voltage, e is the elementary charge, t is the time of flight and d is the length of the drift zone.
Early MALDI instruments were operating based on the linear TOF spectrometer. In the linear mode, ions of different masses meet the detector based on the time that they need to travel across the drift zone. As it is expected, ions of higher weights reach to the detector later than the lighter ions. Therefore, heavier ions occupy higher values of m/z than the lighter ones (Fig.3). In theory, since all the ions receive the same amount of kinetic energy, the ions of same m/z should have the same time of flight and subsequently arrive to their destination (ion detector) at the same time. In practice, however, the exact same intensity of the laser irradiation is not felt by the ions of same m/z. For that reason, there exist a distribution of kinetic energies in each discrete m/z. This phenomenon, to a considerable extent, reduces the resolution of the spectrum by assigning a distribution of m/z. The linear TOF method has high sensitivity and high efficiency, with the ability to analyze molecules. However, a limitation of the linear TOF method is that it provides a poor resolution due to the peak broadening that can occur due to the spatial distribution of analyte molecules on the surface and the unequal distribution of energies from the laser pulse. This results in ions with the same m/z having different kinetic energies. Pulsed-ion extraction (PIE) was designed to resolve limitations associated with peak broadening. In PIE, there is a delay in the application of the acceleration voltage following ionization. Ions that gain more kinetic energy in the ionization process will drift away from the target plate. When the acceleration voltage is applied, there is a gradient between the target plate and ground, and the ions farther from the target will experience lower voltage and will therefore have a lower deposited kinetic energy. This phenomenon results in averaging with ions that received less kinetic energy in the ionization process, thus normalizing the kinetic energy of ions having the same m/z ratio.
The lack of resolution in linear mode was corrected by a more advanced technique, reflection mode. This sophisticated technique consists of ion mirrors and is equipped with electric fields that repulse back the ions into the drift zone and force them to fly back to the detector, which is now installed in the opposite side of the flight path. The reflection mechanism that refocuses the analyte ions by their masses hence results in improved resolution [12].
A reflectors is a focusing element at the end of the TOF instrument that changes the direction of ion travel. A voltage is applied to these lenses and causes a change in the trajectory of that ion. Ions with a higher kinetic energy will penetrate the reflector deeper than those with a lower kinetic energy, such that the flight path is elongated, allowing for averaging of flight times and decreasing peak broadening. Although a reflectron is effective at reducing peak broadening, it essentially doubles the ion path; therefore, when sensitivity is an issue, it is necessary to use linear TOF due to the potential for ion scattering. For this reason, when analyzing high-mass ions with MALDI-TOF MS, as in the case of samples utilizing commercial MALDI-TOF MS platforms, linear TOF is most commonly used [13].
MALDI analysis requires the use of laser beam in different ranges as follows: Ultraviolet (UV) MALDI applies laser beam in the UV range such as nitrogen lasers at 337 nm. UV lasers are the most widely applied laser range in MALDI analysis. IR-MALDI although use of infrared (IR) range of laser beam is not as common as UV, it founds its application as it offers softer mode of ionization. IR-MALDI is also privileged with the greater material removal (in the case of biological applications) and higher compatibility with other mass spectrometry from the category of matrix-free laser desorption techniques [14].
The plate is subsequently loaded into the MALDI-TOF MS instrument and analyzed by software associated with the respective system, allowing rapid identification of the bacteria [9]. The software for the MALDI system contains a database with information about which fragments one can possibly get from a particular bacterium. This information can be used by the system to compare with unknown samples and calculate a score value for each sample. The score value is a measure of the probability that a particular isolate will represent a particular bacterial species/strain in the reference library. The identification criteria of MALDI Biotyper ™ system log (score values) higher than 2.0 indicates species identification, while log (score values) between 1.7 and 2.0 are sufficient for identification of the genus. For a score value between 0.000 and 1.7 means that identification is not possible since the peak of the unknown isolate does not represent any bacterium in the reference library so , no isolate identification is possible [15].
After drying, the mixture of the sample and matrix co-crystallizes and forms a solid deposit of sample embedded into the matrix. The plate is subsequently loaded into the MALDI-TOF instrument and analyzed by software associated with the respective system. The MALDI leads to the sublimation and ionization of both the sample and matrix. These generated ions are separated depending on m/z through a TOF analyzer, and a spectral representation of these ions is generated and analyzed by the MS software, generating an MS profile.
Direct method: In this method, bacteria colony were applied, as thin films on the target plates using sterile tooth picks. The bacterial smear were then left to dry at room temperature for 5 min. Subsequently, 1 µl of the matrix solution, was applied to the samples and co-crystallized with them at room temperature for 10 min. After drying insert the plate in to MALDI TOF MS and then read the result of the samples. .
Extended method: In direct extended method, bacterial colony was applied and dried on the target plate as in the direct method. Following this, 1 µl of 70% formic acid was deposit on the samples on the plate by pipetting. Finally, 1µl of the matrix solution was applied onto the spot dried at room temperature for approximately 10 min. .
Ethanol-Formic Acid Extraction method: A loop-full colonies were transferred using a 5 µl inoculating loop into 300 ml of distilled water and 900 ml of ethanol. The suspension was pelleted after centrifugation at 12,000 rpm for 2 min, dried, and then reconstituted in 20–80 ml of 70% formic acid (select volume based on the pellet size). After incubation for 30 s, equal volume of acetonitrile was added. The suspension was then centrifuged at 12,000 rpm for 2 min. A volume of 1.0 µl of the supernatant was applied to spot target plate .
Trifluoroacetic acid extraction method:Place a loop-full of culture using a 5 µl inoculating loop into a tube and add 50 µl of 80% TFA. Suspend the cell mass by pipetting until complete dissolution and denaturation of the sample achieved. Add 3 volumes of double distilled water (150 µl) and 200 µl of pure acetonitrile and mix by vortexing. Centrifuge at 12,000 rpm at room temperature for 2 min. A volume of 1.0 ml of the supernatant was applied to spot target plate .
Disruption method: Physical disruption method: This procedure is identical to ethanol-formic acid extraction except one additional step that the hydro-alcoholic suspension of sample heated at 95◦C for 1 h before extraction. The step of heating can also be replaced with mechanical lyses by 3 cycles of micro-beads beating (60 s) before extraction as described above in ethanol-formic acid extraction. .
Enzymatic lysis method: Microbial biomass were harvested and washed three times with deionized water. The cell pellet resuspended in 30 µl of water were digested by adding 10 µl of 1 mg/ml lysozyme and incubated for 30 min at 37◦C. Termination of digestion was accomplished by addition of 0.1% TFA to the pellet. .
Bacterial Test Standard mass calibration or Bruker BTS contains a carefully manufactured extract Escherichia coli DH5 alpha that express a characteristic peptide and protein profile plus additional proteins for use in MALDI-TOF-MS. The extract is spiked with two additional proteins that extend the upper boundary of the mass range covered by Bruker BTS. The overall mass range covered by Bruker BTS is 3.6 to 17 kDa. Each tube contains enough material for the preparation of around 40 MALDI target plate spots. Bruker BTS is shipped at ambient temperature but must be stored at -18°C or below immediately on arrival. The expiry date on the package is valid for the enclosed Bruker BTS when stored at -18°C or below on arrival[16].
Bruker BTS must be solubilized before use strongly recommend that dissolved Bruker BTS is aliquoted and frozen using 0.5 mL screw cap micro tubes with screw caps. Frozen, dissolved Bruker BTS can be stored for up to 5 months at –18°C or below. With time, oxidation products may be detected. These oxidation products do not significantly influence spectra quality. FlexControl Software versions 3.0 and earlier require a free software patch to support Bruker BTS.
Bruker BTS Preparation Procedure: Poor sample preparation will degrade sensitivity and will yield low resolution and poor reproducibility. The generation of ions through MALDI depends on the presence of an optimal ratio of matrix substance to analyte. For best results, use freshly prepared solutions and chemicals of the highest purity available.
Bruker Bacterial Test Standard (BTS must be prepared as described). Standard solvent (acetonitrile 50%, water 47.5% and trifluoroacetic acid 2.5%) from Sigma- Aldrich which has been tested by Bruker Daltonik GmbH and is for Solubilization of Bruker BTS
Solubilization and Preparation of Bruker BTS Aliquots: Add 50 µL of standard solvent to the Bruker BTS pellet and dissolve by pipetting up and down at least 20 times at room temperature. Incubate the solution for 5 minutes at room temperature and then mix by pipetting up and down at least 20 times. Centrifuge at 13,000 rpm for 2 minutes at room temperature. Pipette 5 µL aliquots of the supernatant into screw cap micro tubes and close tubes tightly. Store the aliquots at –18°C or below.
Frozen Bruker BTS aliquots should be thawed at room temperature and mixed by flicking the screw cap micro tube with a finger. Screw cap micro tubes should be closed and stored at ─18°C or below immediately after preparation of Bruker BTS sample spots. Each 5 µL aliquot contains enough material for at least fourtheen (40) sample spots [16].
Matrix used in MALDI-TOF MS are generally crystalline solids with low vapor pressure that can easily become volatilized to form ions in a vacuum matrix is mixed in excess with a sample and allows for production of intact, gas phase ions, nonvolatile, and thermally labile compounds such as proteins. The matrix plays a key role by absorbing the laser light energy and causing a small part of the target substrate to vaporize. Matrices should possess certain characteristics, such as having a strong absorbance at laser wavelengths used to facilitate ionization, stability in a vacuum to force an interaction with the co ionized specimen, an ability to ionize the specimen, solubility in solvents that are compatible with the samples in order to create an effective matrix-specimen mixture, and a complete lack of any chemical reactivity with the specimen, to avoid unwanted alterations or damage to peptides contained within the sample. In the case of MALDI-TOF MS, which uses a UV laser, the matrix molecule must also have a strong Chromophores as part of its composition to help absorb energy, thus preserving the protein fragmentation. Chromophores are selected based on their ability to absorb specific laser wavelengths, resulting in electronic excitation of the matrix. Some of the commonly applied matrices in MALDI analysis are: (i) are 3,5-dimethoxy-4-hydroxycinnamic acid (sinapinic acid); (ii) a-cyano-4- hydroxycinnamic acid (CHCA, alpha-cyano or alpha-matrix); and (iii) 2,5- dihydroxybenzoic acid (DHB)(Gross et al., 2006). CHCA is commonly used for peptides in the lower mass range (< 2,500 Da), as it is considered a “hard” matrix, which means that a large amount of internal energy is transferred to the analyte during desorption and ionization, causing a considerable amount of ion fragmentation in the drift tube. The main advantage of CHCA in the analysis of peptides is that it results in the formation of small homogenous crystals, yielding good resolution during MS analysis. SA is most commonly used in the analysis of high mass peptides and proteins (> 2,500 Da). Compared to CHCA, it is a “soft” matrix. When an SA matrix is used, the analyte receives less internal energy deposition, resulting in less analyte fragmentation. Thus, this matrix is more suitable for analysis of proteins. SA can also promote the formation of small crystals. DHB is the matrix of choice for the analysis of glycoproteins and glycans, and it is also often used for peptide analysis. The main disadvantage of DHB is that it forms large crystal needles, which create large variations in sample geometry from spot to spot within a sample. When spectra are summed from different sample preparation spots, the resolution is considerably lower than that of spectra obtained from a CHCA preparation. On a steel target, DHB preparations will form a crystalline ring. Good peptide spectra are usually obtained only from the rim of the preparation. The main advantage of DHB for the MALDI analysis of peptides is that this matrix is more tolerant to contamination (with salts and/or detergents, for example) than other matrices [1].
Bruker Matrix Preparation Procedure: Add 250 µL of standard solvent (acetonitrile 50%, water 47.5% and Trifluoroacetic acid 2.5%) to the Bruker Matrix pellet and dissolve by vortex until free of crystal. Avoid foaming of the solution! Store at room temperature for 5 days..
Preparation of HCCA Bruker Matrix on Samples for MALDI Target: Smear bacteria colony on spot of the MALDI target. Dry the samples at room temperature. Then overlay the 1 µL HCCA Bruker Matrix solution on sample spot. Dry the matrix at room temperature. Deposit 1 µL Bruker BTS solution immediately after drying. Dry the Bruker BTS at room temperature. A homogeneous preparation should be observed [16].
One of the most important steps for MALDI-TOF MS analysis is choosing the proper matrix for the analyte molecules of interest. The most commonly used matrices for biological samples, including proteins and peptides. A suitable matrix consists of crystal molecules that, typically, in association with ionizing agent allows the analyte to be co-crystallized in a homogeneous modus. When a mixture of matrix and analyte is mounted on the MALDI plate, Reflection mode in MALDI-ToF-MS and application of mirror method in correcting the error produced by linear mode. Principles and Mechanism of MALDI-ToF-MS Analysis evaporate leaving behind the crystallized analyte along with the recrystallized matrix in the MALDI spot. This phenomenon is called cocrystallization. The quality of the co-crystallization is of a major importance in matrix selection. Desirable matrix should possess:Specific range of molecular weight that permits easy evaporation of the matrix but not in the sample preparation process. Preferable acidic nature for the matrix that acts as the proton source and promotes the ionization. Wide range of optical absorbance that covers either UV or IR range. Functionalization with polar groups and subsequent applicability in aqueous medium [17].
Variety of matrix options has been investigated in MALDI analysis and different levels of efficiencies. Many of the researchers, however, reported frequent incompatibilities, background effects and limitations for the used matrices. Particular matrices in association with specific ionizing agents may offer limited stability that can last only for few minutes. Obscurity in mass spectrum might occur as some of the matrices show self-polymerization when mixed with analyte and ionizing agent. Specific matrices have proven to lead to a significant level of mass discrimination. Liquid matrices in special cases might cause intense contamination of the ion source. In peptides analysis, matrices with low protein affinity (PA) values have induced more fragmentation than matrices with high PA values [18].
Specialized applications such as identification of microorganisms using MALDI Biotyper may require use of a specific MALDI target type. Steel targets: Standard target for fast, simple and robust MALDI preparation of virtually any type of sample. Two types of steel target are available [5].
Ground steel targets have a highly regular fine structure on the plate surface, enabling highly homogenous co-crystallized preparations (dried droplet method).
Polished steel targets are virtuallly free of any surface structure, and provide maximum flatness for MALDI thin-layer preparations.
AnchorChip Standard targets is target type for high-throughput MALDI measurements that are performed in unattended, fully automatic mode. Such applications include MALDI peptide mapping and subsequent MS/MS sequencing of 2D gel digests and LCMALDI analyses of complex peptide mixtures.
Sample positions on Anchor Chip targets contain "anchors"; hydrophilic patches surrounded by a hydrophobic ring. The "anchor" localizes droplets at the sample position and the hydrophobic ring prevents sample spreading and concentrates the sample into a spot 800 µm in diameter. After correct adjustment of the target in the MALDI ion source, the localization effect ensures that every single laser shot fired throughout an automatic run will hit a sample spot. This significantly increases the efficiency of the MALDI acquisition process. The concentration effect also provides enhanced sensitivity when analyzing dilute samples
Small Anchor target: Preferred target type for the preparation of oligonucleotides and similar samples using 3-HPA as a matrix
Prespotted AnchorChip (PAC) targets: Disposable MTP-sized MALDI targets prespotted with HCCA (a-cyano-4-hydroxycinnamic acid) matrix. PAC targets are highly suitable for low- to medium- complexity samples in gel- or LC-based proteomics applications [5].
MALDI Biotyper RTC: (Real time Classification) aimed at users who want to classify unknown samples quickly and easily. Users simply load samples, start the classification and wait for a result.
MALDI Biotyper OC: (Offline Classification) aimed at advanced users who want to fine-tune the classification process and investigate the relationships between groups of organisms
Flex Control: Used to control the MALDI- TOF instrument. Can run in hidden mode during MALDI Biotyper classification runs.
Currently, biochemical tests form the basis of most techniques used to identify microorganisms. Reagents that are metabolized only by specific organisms are added to cultures. This leads, for example, to a characteristic color change of the added reagent, which can then be compared with reference values. Correlating the results of a number of such tests on a sample culture provides an indication of the organism's identity [19]. The MALDI Biotyper Real time Classification User Interface window contains a title bar, menu bar, toolbars and a status bar; MALDI target and legend displays; anMSP view, and a results table [19].
MALDI Biotyper Offline Classification (OC) uses the same database and offers the same classification algorithms as MALDI Biotyper Real Time Classification (RTC) but gives users more flexibility in defining classification parameters. MALDI Biotyper OC software also enables users to optimize the classification process by creating, modifying and organizing reference spectrum data. Advanced analysis and output options enable visualization of relationships between different groups of microorganisms and test samples using Principal Component Analysis (PCA) and Composite Correlation Index (CCI) methods.
Flex Control is a program designed to configure and operate time- of- flight mass spectrometers of the Bruker micro flex, auto flex and ultraflex series. Flex Control runs under Microsoft Windows 7 (SP1) or XP (SP3) operating s;ystems. Both manual and automatic calibration procedures are available. The manual procedure can be used with all versions of flexControl whereas for the automatic procedure flex Control 3.3 or later is required
Among approaches to identify bacteria with mass spectrometry, MALDI TOF MS is most commonly used, due to its ease of use and the speed with which data can be collected. Most common, a library-based strategy is employed in which spectra of unknown bacteria are compared to libraries that contain spectra of known, reference bacteria. As with conventional methods of microbial identification, MALDI TOF MS enabled identification nearly always requires Cultivation of the microorganism of interest. Solid agar media, liquid broth media have both been used. Cultured bacterial cells are prepared for MALDI analysis. Either intact cells or cell extracts are prepared for analysis. With intact cells, MALDI matrix (e.g., ferulic acid or sinapinic acid) is mixed directly with intact cells. With cell extracts, bacterial cells are lysed via physical (e.g., sonication or bead-beating) or chemical (e.g., via exposure to TFA and formic acid/organic :
solvents) means to release the contents of the cells into the supernatant. The supernatant is added to matrix and analyzed with MALDI.
A variety of mass spectrometers that range from relatively low-resolution, linear detector-only benchtop units (e.g., Bruker’s Microflex LT) to units with much higher resolution (e.g., Shimadzu’s Axima series), can rapidly generate spectra from intact bacteria and cell extracts. Spectra are typically collected with the linear detector of MALDI TOF MS units but a few investigators have used the reflector detector higher resolution units) to obtain higher resolution.
Data. Spectra are typically collected from 2 to 20 kDa although broader and narrower ranges have both been used with success. Peaks in spectra are not typically identified in library-based approaches. While it has been hypothesized that peaks represent proteins, particularly those of ribosomal origin [20].
MALDI Biotyper analysis generates a characteristic mass and intensity distribution of the mainly ribosomal proteins. Because this mass spectrum is species-specific for a large number of microorganisms, it represents a 'molecular fingerprint'. Unknown microorganisms can be identified by comparing their 'fingerprint' with the thousands of patterns in the reference database. The first step of data processing is transferring acquired mass spectra into peak lists. In a second step, these peak lists are compared with every pattern in the reference database [21].
A sample of the unknown micro-organism is applied onto a polished steel plate and allowed to dry. This sample may be whole cells (directly transferred from culture) or protein extracts. A matrix solution, which is made up of the matrix (generally a crystalline solid), aqueous acid and organic solvents, is then applied to the sample. During the drying of the matrix solution there is disruption of the cells (if a whole cell sample) and crystallization of the sample and the matrix resulting in the sample embedded within the matrix.
The steel plate is then inserted into the mass spectrometer and a laser is fired in short bursts resulting in vaporization and desorption of the sample and matrix from the metal plate. In addition to its scaffolding function, the matrix is also important for absorption of the high laser energy and providing the protons for ionization of the sample proteins. An acceleration voltage is applied to the ionized particles, which then enter a flight tube with a detector placed at the other end. This detector analyses the time of flight of the ions through the flight tube. These charged particles are separated by their mass/charge (m/z) ratio, with smaller proteins travelling more quickly than larger proteins. MALDI generally results in singly charged particles and consequently a spectrum is generated, with peaks representing different masses and the height of the peak representing the intensity of the signal [21].
The raw mass spectrum generated requires further data processing before the proteomic fingerprint can be compared with the spectra in the reference database. The comparison algorithms and databases used may vary between different systems but in general the principle is similar. A score is generated depending on the number of matching peaks and a list of top matches together with the probability of the identification (from highly probable species identification to no reliable identification)isproduced [22].
The MALDI Biotyper uses a different methods to identify microorganisms by analyzing samples proteins. Mass spectral pattern of protein is compared with reference patterns in a database. Starting material for identification using the MALDI Biotyper is an individual colony from an overnight culture on an agar plate. The sample is transferred to a position on a MALDI target and air-dried. After drying, a small quantity of matrix solution is added. The organic solvent in the matrix solution extracts proteins from the microorganisms. The extracted proteins are mainly ribosomal proteins, which are present in high concentrations. Once the matrix has crystallized, sample preparation is complete and the samples can be analyses. If this straightforward sample preparation method is unsuccessful, a supplementary protocol containing short chemical extraction steps can be used to increase the range of organisms that can be identified by MALDI analysis [23].
A laser in the MALDI-TOF mass spectrometer irradiates the matrix/sample mixture, evaporating the matrix and releasing positively charged proteins in a so-called “soft” ionization process. The ability of the matrix to absorb UV light and transfer protons onto the extracted proteins is crucial to this process. The protein ions are electrostatically accelerated over a short distance and arrive in the flight tube at a speed that is proportional to their mass. Protein ions with different masses arrive at the detector after different time periods. Simply by measuring the time between pulsed acceleration and the corresponding detector signal (in the nanosecond range), the speed of the ions can be measured very precisely and converted into an accurate molecular mass [9].
There are several commercial vendors that can identify bacteria and yeast using MALDI-TOF MS. Most of these databases may be enlarged to contain spectral entries that are not present in commercially available versions. Further selective analysis utilizing MALDI-TOF MS, such as strain typing and epidemiological research, requires the capacity to add spectra and create customized databases. Each proprietary system has its own set of software, databases, algorithms, and interpretative standards for identifying microbes [23].
Any time a new diagnostic technique is developed, its repeatability, accuracy, and robustness should be assessed against existing procedures that serve as reference standards. MALDITOF MS accuracy compared to Vitek 2, and API-based procedures, as well as phenotypic and molecular approaches. MALDI-TOF MS for the identification of certain microbe groups, as well as to investigate the technology's potential and applications for specialized testing relating to particular microbiological genera [24].
There are several pathogenic bacterial species that belong to the gram-positive bacterial that are found in the microbiology lab. The cell wall of these bacteria, in general, contains a lot of peptidoglycan, which is employed to display a variety of surface proteins involved in adhesion to and interaction with host tissues. Numerous pretreatment or enhancement techniques were developed to address this issue, including the addition of formic acid, lysostaphin, lysozyme, mutanolysin, and proteinase K to bacterial suspensions. This thick layer of peptidoglycan can occasionally make these bacteria more resistant to lysis than their Gram-negative counterparts [25].
Gram-positive isolates are prepared for MALDI-TOF MS detection. Before analysis, Gram-positive organism samples were processed using entire cells and a protein extraction step. The approach that protein extraction with a formic acid (FA) overlay. The amount of unidentified isolates rose with repeated passage of organisms, although incubation temperature and medium type (blood, chocolate, etc.) had no impact on MALDI-TOF MS identification.
Protein extraction is frequently performed prior to MALDI-TOF MS analysis to increase the method's ability to detect Gram-positive organisms. Additionally, even though commercial software databases were suitable for diagnostic use, this resulted in the development of a collection of reference spectra and helped microbiology laboratory staff members learn how to create spectral collections internally for the identification of uncommon species that weren't yet included in commercial databases [26].
Gram-negative bacteria are encountered in the bacteriology laboratory in all sample types analyzed and are ubiquitous members of the normal flora. For example Enterobacteriaceae represent a dynamic group of organisms encountered in the laboratory. This broad group of organisms is often characterized by the ability of the organisms to ferment lactose, as determined via biochemical testing. MALDI-TOF MS was utilized by investigators to identify members of this large group of bacteria and classify them as such while the technique was still in its infancy. Prior to the advent of comprehensive databases that can be easily queried with acquired spectra, early work using MALDITOF MS for the identification of the Enterobacteriaceae was focused on the identification of suitable biomarkers for group-level and genus-level identification. One early study from 1999 using intact cells lysed by one cycle of freeze-thaw from storage at 20°C examined the use of MALDI-TOF MS to distinguish between members of this group of organisms: E. coli O157:H7, Klebsiella pneumoniae, Salmonella enterica and Providencia rettgeri [27].
With respect to sample processing for accurate identification and the analysis of variables potentially influencing MALDI-TOF MS identification. Gram-negative identification using the Biotyper system. Gram-negative organisms, the organisms were spotted onto a MALDI plate using either a light or heavy smear (correlating to the amount of inoculum) or either overlaid with 1 μl of formic acid or not overlaid. Samples were finally mixed with matrix solution and analyzed by using the Biotyper 3.0 software (Bruker) [28].
Identification of Mycobacterium tuberculosis and other mycobacteria from clinical specimens often requires a number of diverse techniques for observation, recovery, growth, and characterization. These closely related strains aside, the rest of the isolates could be easily and accurately identified to the species level, demonstrating the utility of MALDI-TOF MS for the identification of mycobacteria in the microbiology laboratory. The implementation of MALDI-TOF MS in the laboratory will provide a powerful and accurate tool to quickly identify mycobacteria from culture. This implementation will also change testing algorithms for mycobacteria and provide a mechanism for enhanced surveillance and epidemiological data worldwide. The implementation of MALDITOF MS technology will reduce testing costs for mycobacterial identification, with consumable costs per specimen estimated to be less than $1 per isolate [29].
Procedure representing a synthesis of inactivation and processing methods. Colonies were collected in screw-cap tubes containing water and 0.5% Tween 20 and inactivated by heating at 95°C for 1 h. Inactivated samples were centrifuged, washed twice with water, and then vortexed with glass beads to facilitate complete cellular disruption. Following centrifugation, the pellet was resuspended in FA-acetonitrile and centrifuged again. Finally, the supernatant was deposited onto the MALDI test plate and overlaid with matrix [30].
The MALDI-TOF MS has special method in routine identification of pathogens that require long incubation times for isolation and are biochemically inactive, such as anaerobic bacteria. MALDI-TOF MS has also been utilized to evaluate subsets of anaerobic microorganisms isolated from defined biological niches. The applicability of this method for routine identification of important an imal and human-pathogenic anaerobic bacteria. Culture specimens were subcultured and grown in an anaerobic chamber [31]. MALDI-TOF MS for the identifications of anaerobic bacteria, with MS technologies likely to be implemented as the primary determinative method in the future as the technology continues to develop. It is also important to emphasize that the ability of MALDITOF MS to identify anaerobic species currently is not as robust as it is for the routine species-level identification of other groups of bacteria; therefore, the use of additional confirmatory testing will likely be necessary.
Specific protocols for sample preparation for the analysis of anaerobic bacteria have also been investigated. Three pre analytical processing methods for MALDI-TOF MS analysis of anaerobic bacteria: direct smear of intact cells,chemical extraction, FA extraction for MALDI-TOF MS identification of anaerobic bacteria. Bacterial isolates were smeared directly onto the MALDI plate, with 70% FA being directly deposited onto the smear. The samples were subsequently overlaid with matrix and analyzed by MALDITOF MS. Samples were analyzed with the Bruker Biotyper version 3.0 software, with manufacturer-recommended cutoff log score values being used for genus- and species-level determinations [32].
Delayed and incorrect diagnoses are potential risk factors leading to high mortality rates due to invasive aspergillosis and other systemic fungal infections. Because of variability of growth patterns and difficulty obtaining a standardized inoculum, mold identification proves to be more difficult than identification of bacteria and yeast. Phenotypic identification of molds requires experienced and skilled mycologists, who are not available to all laboratories, while molecular methods experience limitations with respect to difficulties associated with lysis and PCR inhibition of mold specimens. Moreover, phenotypic identification is slow and labor-intensive, with many species being phenotypically similar but genotypically distinct or different in their propensity to cause disease. Therefore, a rapid mechanism requiring minimal sample preparation and analysis by the specialist for the identification of these organisms is appealing. MALDI-TOF MS has been evaluated to potentially fill this niche. A number of studies have examined the ability of MALDI-TOF MS to identify fungi. Implementation of the database has improved laboratory efficiency and turnaround time [33].
Sample preparation methods for MALDI-TOF MS analysis of fungal hyphae and spores have been identified by five methods for the preparation of samples from this material, using various combinations of culture conditions (agar versus broth), ethanol and no ethanol treatments, heat treatments, and mechanical lysis procedures to determine the method [34] . Fungi were cultivated on a Sabouraud gentamicin-chloramphenicol agar plate for 72 h at 27°C and then extracted with FA following incubation in ethanol. Acetonitrile was added, and the mixture was centrifuged, with the resulting supernatant being analyzed by MALDI-TOF MS. Mechanical lysis method for the preparation of samples from mold specimens for MALDI-TOF MS analysis. Small piece of mold isolate was resuspended in ethanol and zirconia-silica beads, emulsified thoroughly, and vortexed. Following centrifugation, the supernatant was removed, and the remaining pellet was resuspended in 70% FA, vortexed a second time, and centrifuged. The resulting supernatant was then either analyzed immediately or stored for up to 1 week at 20°C for subsequent analysis by MALDI-TOF MS. Both methods worked well for the investigators in protocols will add value toward the future standardization of sample preparation methods for fungi specimens. Irrespective of which methods is chosen in the future, the use of a biosafety cabinet during sample processing and the minimization of aerosols will be important considerations during protocol selection to ensure the safety of laboratory personnel working in close proximity [35].
An appropriate drug regimen with antifungal treatment is a critical component to successfully cure serious fungal infections, but targeted therapy is difficult. It requires several days for growth and identification by conventional biochemical and morphological approaches, and the results may sometimes be inconclusive. Species-specific susceptibility patterns can help clinicians make therapeutic decisions, but determination of antifungal susceptibility can take weeks in some cases. Protein content and expression levels of fungal isolates may be affected by growth states and mycological life cycles; therefore, standardization of medium and growth phase will be important when using MS technology for evaluation of therapeutics. Still, MALDI-TOF MS holds the promise of significantly accelerating these processes, substantially improving fungal diagnostics and patient treatment.
The rapid analysis of yeast by MALDI-TOF MS has the potential to revolutionize. Identification of yeast in the laboratory is most traditionally associated with the use of selective or differential media in addition to both manual and automated biochemical identification systems. By implementing a rapid system which functions on a universal platform for the definitive identification of yeast specimens, significant reductions in time to diagnosis and laboratory costs could be realized.
Yeast isolates were analyzed by directly smearing a small amount of biomass onto a MALDI plate and overlaying the smear with 70% FA, followed by drying and addition of matrix. MALDI-TOF MS analysis was performed with the Bruker BioTyper 3.0 database, and a log score of 1.7 or higher was accepted for a species-level identification [3].
In addition to the accurate identification of bacteria in sample another critical responsibility of the microbiology laboratory is to determine antibiotic susceptibilities of organisms and to report these findings to for the implementation of targeted antimicrobial therapy. Mechanisms used for the determination of antimicrobial susceptibilities can vary depending upon the type of organism in question but can include culture-based methods such as Kirby-Bauer disc diffusion testing, Etests, selective growth media, and broth micro dilution analysis. Additionally, many automated technologies used for the phenotypic identification of bacteria, including the Vitek-2 (bioMérieux) and the Phoenix (Becton, Dickinson) systems, also function to determine antimicrobial susceptibilities. The drawback of these technologies is the increased time to result associated with them. Rapid testing mechanisms for commonly encountered resistant organisms (i.e., MRSA) were developed as well, but there are limited to a small number of bacterial species. What is not currently available is a universal platform for the rapid determination of antimicrobial resistance covering an extended spectrum of bacterial genera that can be implemented into the workflow of the laboratory [36].
Complications that arises when identifying organisms based on traditional or molecular methods is that it can be difficult to discriminate among species that are phenotypically, biochemically, or even genetically similar. Depending on the similar species organism, are grouped together (e.g. coagulase negative staphylococci) or an incorrect identification at the species level. MALDITOF MS, depend on the measuring bacteria proteins that are typically well conserved within a species. This is a more reliable means of discriminating one species from another. MALDI-TOF MS is able to distinguish between closely related bacterial species with a high degree of confidence [37]. This is especially beneficial for organisms in which an incorrect identification or lack of a species level identification. This could include species that are predictably resistant to specific antibiotics, those that have limited therapeutic options [38].
Among the organisms that are particularly difficult to identify to the species level using traditional methods, but readily identified by MALDI-TOF MS. For example, one study found that MALDI-TOF MS correctly identified more than 86% of HACEK (Haemophilus, Aggregatibacter, Cardiobacterium, Eikenella, Kingella) isolates, whereas biochemical testing identified less than 77% [39]. A substantial cost savings can be achieved by implementing MALDI-TOF MS as the primary method of identification in the diagnostic laboratory. While the initial cost of the instrument is high, the cost savings on reagents and labor can offset the expenditure.
Reagent savings will vary from one laboratory to another, but for the typical moderate to high volume lab, a savings of greater than 50% would be anticipated. This technology improves more species are identification and additional cost saving [40] . Significantly decreases the turnaround time, sample preparation is simple and the sample requirement is minimal and single colony is sufficient in order to generate spectra of sufficient quality. Cost effective low consumable costs, automated, robust, interlaboratory reproducibility, broad applicability (all types of bacteria including anaerobes, fungi) [41].
MALDI TOF MS in the diagnostic laboratory can accurately identify most closely related species. However, there are some exceptions. The inability to discriminate between related species can be due to the inherent similarity of the bacteria themselves. For example, MALDI-TOF MS is currently unable to differentiate E. coli from Shigella [37]. This is likely because these may not be two species, but actually one, as has been suggested by taxonomists. In the future, the addition of proteomic based approaches to the typical MALDI-TOF MS system may improve the discriminatory power of this method and make it possible to identify organisms at the strain or serotype level. Another reason similar species may be incorrectly identified is due a lack of sufficient spectra in the database [42].
Identification of new isolates is possible only if the spectral database contains peptide mass fingerprints of the type strains of specific genera/species/subspecies/strain. No susceptibility information is provided, not useful for direct testing of samples (except urine and blood). Some organisms require repeat analysis and additional processing (extraction). The acceptable score cutoffs vary between studies and some closely related organisms are not differentiated [21]. The important challenges in species identification arise from incomplete databases, close relatedness of species of interest, and spectral quality, which is currently vaguely [43].
The usefulness of MALDI-TOF MS, which is currently widely used in diagnostic microbiology laboratories, is being further investigated. Specifically, to increase the quantity of microorganisms discovered, to recognize microorganisms directly from specimens, to find antibiotic resistance, and to support epidemiological research. It will be necessary to expand reference data banks, standardize identification techniques, and validate them in order to increase the variety of microorganisms that are detected, including the routine identification of antibiotic resistance, bacteria, mycobacteria, and fungi [44]. However, there are still difficulties to be overcome if reliable identification of microorganisms directly from other samples, such as blood, urine, and cerebrospinal fluid is to be achieved. From an epidemiological point of view, the development of MALDI-TOF MS for strain typing, which is still in its infancy, would be a significant boon for infection control and public health [45]. MALDI analysis has rapidly found variety of applications in many different scientific areas. In particular, MALDI analysis provides essential information about disease diagnosis, antibiotic sensitivity, measurement of the additives and impurities, product formulations and disease epidemiology
Main applications of MALDI-ToF-MS in some important areas of science
Disease diagnosis: In the area of microbiology, MALDI-TOF-MS can be applied for identification of microorganisms such as bacteria, mycobacteria and fungi. This technique has a great potential for species identification in the field of veterinary, medical and environmental microbiology.
Antimicrobial sensitivity: MALDI-TOF-MS, in addition to diseases diagnosis is often utilized for antimicrobial sensitivity. Due to the rapid identification of proteins, MALDI holds a great potential in medical and diagnostic applications. Since MALDI-ToF-MS can reveal drug resistance to conventional antibiotics, it can also be helpful in determining the right course of treatment.
EPIDEMIOLOGY: The ability of MALDI-TOF MS to give vital epidemiological data cannot be overstated. Rapid and accurate disease detection by MALDITOF MS will be what will largely draw laboratories to mass spectrometry-based analysis methods. Finding strain-specific information for a representative taxonomy of isolates in outbreak circumstances is one of the major issues encountered by clinical laboratory scientists (CLSs), infection prevention practitioners, doctors, and public health laboratories. Microbial typing is extremely difficult in some circumstances (e.g., Salmonella spp. and Streptococcus spp.) and frequently necessitates time-consuming second-stage testing in order to correctly identify serotype, subspecies, or other taxonomic classifications. When the precise ramifications of a serotype or subspecies in the context of an infection are unclear, this challenge may become even more difficult. Due to their intricacy, many strain typing techniques call either extra instruments or materials as well as a specialized laboratory test method. As a result, not all laboratories routinely undertake these tests, which causes more "send-out" testing and delays in the investigation of diseases and outbreaks linked to hospitals. Delays in detection, the loss of crucial epidemiological data, higher testing costs, and perhaps unreliable or inconsistent results are all possible outcomes of this.
Nanotechnology: In the recent years, MALDI TOF MS has achieved a significant position in the field of nanotechnology. Modern targeted medicine and nanotechnology have given biopharmaceuticals, such as protein and peptide medications, a greater role as therapeutic agents. The use of polymers in the creation of innovative drug delivery systems for regulated or prolonged drug release has opened up new opportunities for MALDI analysis in nanotechnology.
Chemistry: MALDI TOF MS is frequently used in the field of biochemistry to quickly identify proteins and Peptide mass fingerprinting. Catenanes, rotaxanes, dendrimers, and hyper-branched polymers are a few examples of the organic macromolecules in chemistry that have extremely large molecular weights and it is challenging to create molecular ions from such microstructures, many ionization techniques are typically limited in their ability to analyze such macromolecules. Using MALDI analysis molecular weight and chemical structure.
MALDI-TOF MS has already become an integral tool in the diagnostic microbiology laboratory for microbial identification, resulting in faster accurate identification of microorganisms, which allows earlier treatment decisions to be made. Ongoing development of this technology and integration with more traditional and molecular microbiological techniques promises to improve the services offered by the laboratory. Although some of the methods described here are currently still restricted to diagnosis, research and large reference laboratories, the potential of these testing algorithms to move into routine microbiology and public health laboratories seems imminent. The implementation of MALDI-TOF MS in the routine diagnosis will provide a powerful and accurate tool to quickly identify bacteria, mycobacteria, fungi from culture. Improvements to spectral databases and analysis software should optimize the use of MALDI-TOF methods and should reduce the turnaround time for identification of nearly all microbes. In the future, mass spectrometers will be linked with automated antimicrobial susceptibility systems, thus allowing partial or complete automation of routine microbial characterization. The first generation of this automation, the BD Kiestra system, is available from Becton Dickinson. Integration of MALDI-TOF analysis should greatly impact the current processes used in microbiology laboratories. Additionally, research dedicated to adapting MALDI-TOF MS technology result in shorter hospital stays, improved prognoses, and decreased financial burden. Rapid and accurate identification of microorganisms directly from clinical specimens is becoming an expectation for clinical laboratories, essential for optimal diagnosis and treatment of patients with infections, and MALDI-TOF technology may support these efforts in the future, but direct-from-specimen testing methods are not currently available. As the number of studies examining the accuracy and applicability of MALDI-TOF MS in routine clinical situations continues to grow, so too will publications examining the financial and clinical benefits of implementing this rapid technology. MALDITOF MS in identification reduced time and laboratory costs. This review should serve as a model for future investigations regarding the implementation of MALDI-TOF MS in different laboratories and evaluations of associated benefits and drawbacks. Newer determinative technologies are constantly evolving for microbiology laboratories, aiding the evolution of the microbiology laboratory from one of the slowest of all laboratory services to one that is a plastic and dynamic entity able to directly influence patient outcomes with speed and robust accuracy. As we usher in an era of improved response and higher expectation of accuracy, clinical laboratories are consistently striving to extend the capabilities of these new methods, often in partnership with developmental scientists, resulting in novel technologies, such as MALDI-TOF MS, which should shape and define the diagnostic landscape for years.
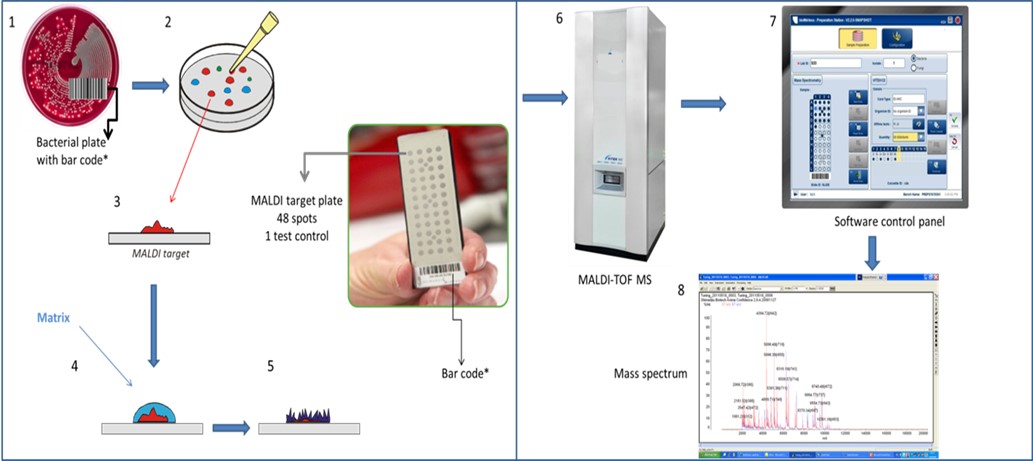 |
| Figure 1: Workflow for microbial identification using MALDI-TOF MS |
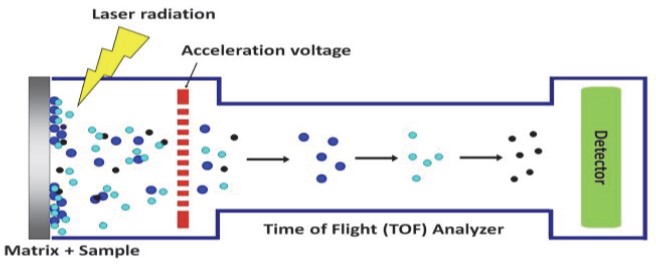 |
| Figure 2: Basic principles of MALDI-TOF MS |
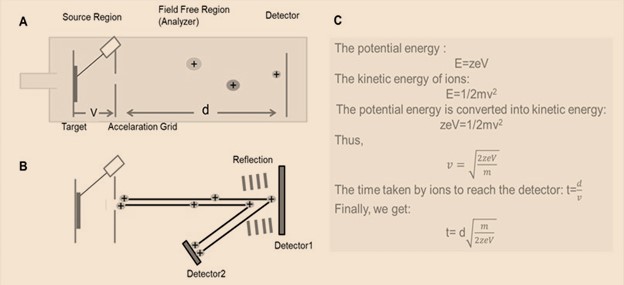 |
| Figure 3: General schematic for TOF analyzer. (A) Liner TOF analyzer; (B) Reflector TOF analyzer; (C) The derivation process of the time that ions pass through field free region in the liner TOF analyzer |
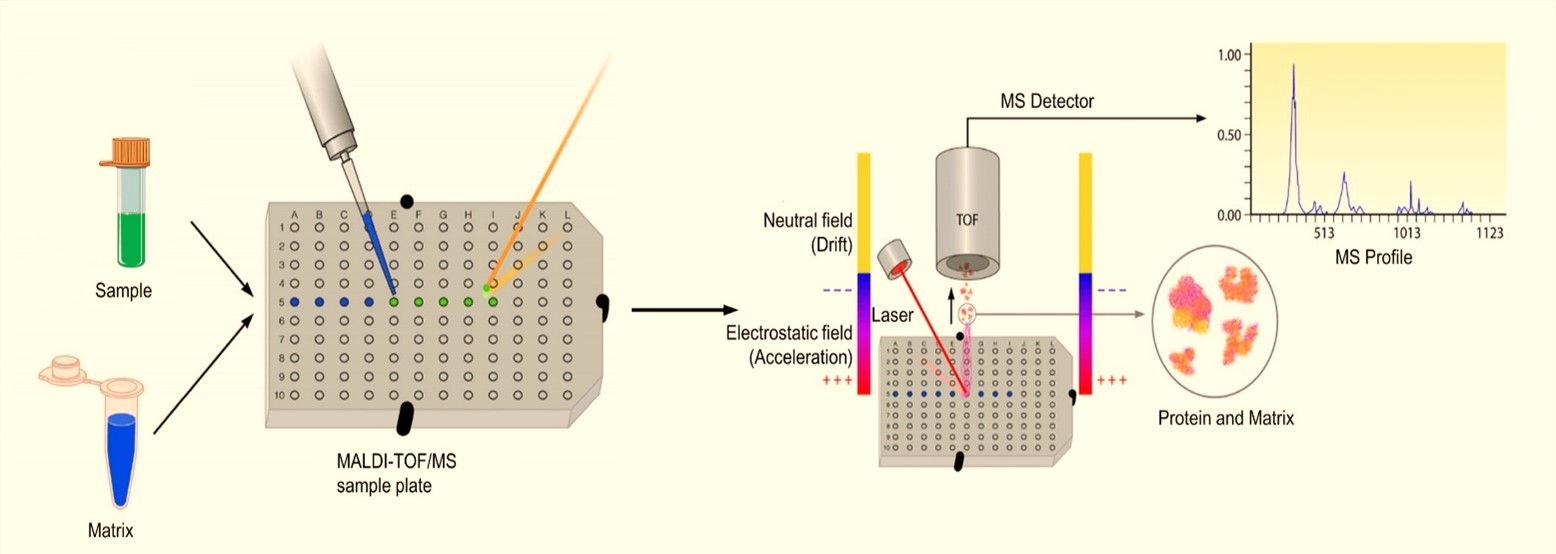 |
| Figure 4: The process of MALDI-TOF mass spectrometry [2] |
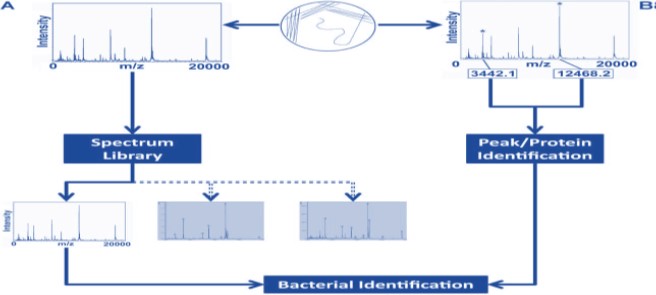 |
| Figure 5: Overview of library-based (A) and bioinformatics-enabled (B) approaches to profiling bacteria using MALDI-MS |






































